Explore our Research Centres
Delivering world class research
Research at the University of Newcastle is supported by a collaborative network of institutes and centres.
Filter centres
University Institutes
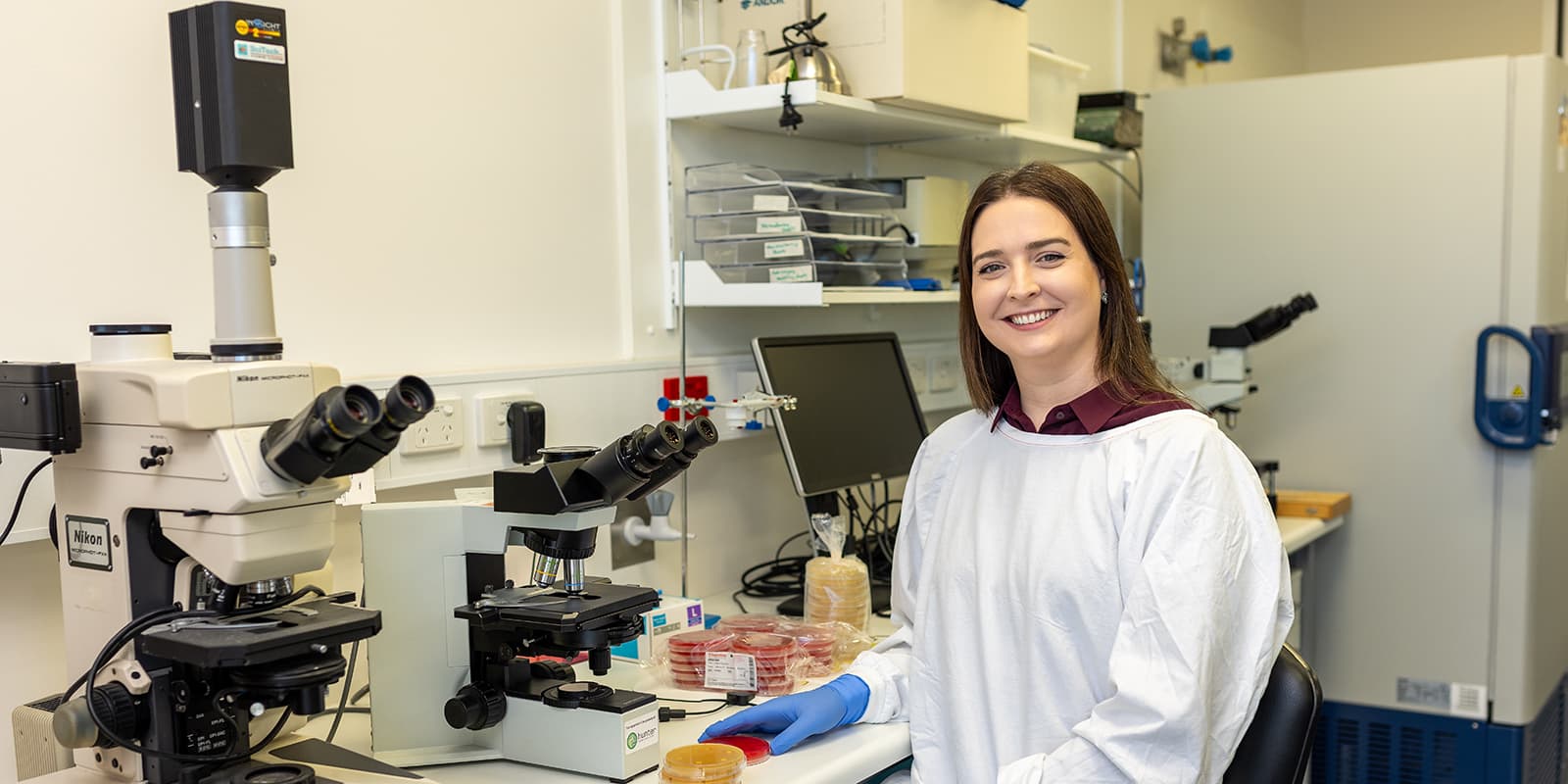
Hunter Medical Research Institute
The Hunter Medical Research Institute (HMRI) supports the Hunter's internationally recognised health and medical research, education and training. HMRI is a multidisciplinary partnership between the University of Newcastle and Hunter New England Local Health District and the community. Established in 1998, HMRI facilitates collaborations between researchers translating scientific advances into better clinical care, competitive commercial products and improved health care guidelines.
Newcastle Institute for Energy and Resources
The Newcastle Institute for Energy and Resources (NIER) is dedicated to addressing global challenges through innovative, multidisciplinary research and strong industry collaboration. Our focus spans critical sectors including energy, resources, food, and water, aiming to enhance environmental, social, and economic outcomes.
Australian Research Council (ARC) centres
Centre of Excellence for Enabling Eco-Efficient Beneficiation of Minerals
This Centre will transform the minerals industry, establishing a new generation of research leaders to support the innovation needed in creating a green economy for future generations.
National Health and Medical Research Council (NHMRC) centres
Centre of Research Excellence in Asthma Treatable Traits
The NHMRC Centre of Research Excellence in Asthma Treatable Traits (CREATT) aims to revolutionise the management of chronic airway diseases by testing and implementing a new paradigm for personalised medicine.
Building on outcomes of the Centre of Excellence in Severe Asthma and its ground-breaking toolkit, CREATT is a consortium of world-renowned clinicians, researchers and respiratory organisations who are generating new knowledge to support the treatable traits approach and ensure translation into practice.
Centre of Research in Digestive Health
The NHMRC Centre for Research Excellence in Digestive Health aims to improve quality of life for patients with unexplained chronic gastrointestinal disorders. It brings together clinical researchers from universities, hospitals and research institutions across Australia and beyond.
The centre supports research, training, multidisciplinary collaboration, and translation of findings to enhance our understanding, identification and management of chronic digestive diseases.
National Centre of Implementation Science
The centre brings together a multi-disciplinary team of emerging and established researchers to generate evidence that helps fast-track the adoption of interventions for chronic disease prevention.
Partner organisations for this NHMRC Centre for Research Excellence include the University of Sydney, Ottawa Hospital Research Institute, Monash University, Central Queensland University and Neuroscience Research Australia.
Cooperative Research Centres (CRC)
Hosted Cooperative Research Centres (CRC)
Soil CRC
The Cooperative Research Centre for High Performance Soils (Soil CRC) is bringing together scientists, industry and farmers to find practical solutions for Australia’s underperforming soils. The Soil CRC aims to enable farmers to increase their productivity and profitability by providing them with knowledge and tools to improve the performance of their soils. It is the biggest collaborative soil research effort in Australia’s history, with 39 Participants that contribute to the Soil CRC through both cash and in-kind contributions.
Partner Cooperative Research Centres (CRC)
HILT CRC
This Cooperative Research Centre aims to help Australia’s heavy industry sector compete in the low-carbon global economy by developing solutions to help reduce its greenhouse gas emissions.
The sector processes minerals such as iron, steel, and aluminium which support the production of critical materials for day-to-day life and makes a significant contribution to Australia’s industrial sovereignty. The HILT CRC will focus on overcoming barriers to the sector’s low-carbon transition.
iMove CRC
The Australian Transport Cooperative Research Centre, known as the iMOVE CRC, is a national transport research and development centre dedicated to providing a coordinated approach to Australian transport research.
It is a consortium of more than 40 industry, government and research partners engaged in a 10-year effort to improve Australia’s transport systems. It aims to deliver solutions that help reduce road congestion, fuel use, emissions, accidents and fatalities. It also works to improve freight co-ordination, productivity, international competitiveness, and overall lifestyle.
MinEx CRC
The MinEx Cooperative Research Centre is working to develop more productive, safer and more environmentally friendly drilling methods for critical deposits that lie beneath sand, soil and sediment. It is also investing the development of new exploration tools and new ways to deploy those tools, to support longer-term sustainability and safety in the sector.
SmartCrete CRC
Concrete is the second most used material on earth after water and is a fundamental element of our built environment. The SmartCrete Cooperative Research Centre provides a nationally coordinated and collaborative platform for research and development in Australia.
It brings together leading industry partners, research institutions, agencies and associations to focus on innovation in engineered solutions, asset management and sustainability.
Research Centres
College of Engineering, Science and Environment
BHP Centre for Sustainable Steelmaking Research
With the global shift to low-carbon iron and steelmaking technologies, the BHP Centre for Sustainable Steelmaking Research is at the forefront of innovation in low-carbon cokemaking, modified blast furnace and alternative iron and steelmaking processes.
Centre for Applied and Responsible AI
The Centre for Applied and Responsible AI (CARA) applies a deep expertise across AI, human-computer interaction, optimisation, data science, robotics, cybersecurity, and digital health.
Centre for Bulk Solids and Particulate Technologies
The Centre for Bulk Solids and Particulate Technologies (CBSPT) is actively involved in both fundamental and applied research on a range of problems associated with bulk solids and particulate technology. Research areas include storage, flow, processing and transportation of bulk solids. CBSPT also provides specialist courses for industry through its professional development programs.
Centre for Conservation Science
The Centre for Conservation Science brings together people like conservation biologists, social scientists, health workers, artists, and communication experts. By working together, they create conservation solutions that can help protect nature here and around the world.
Centre for Construction Safety and Well-being
The Centre for Construction Safety and Well-being (CCSW) is a collaboration between the University of Newcastle and construction company Hansen Yuncken, located within the university's School of Architecture and Built Environment.
Centre for Critical Minerals and Urban Mining
The Centre for Critical Minerals and Urban Mining is concerned with the science and engineering of particulate systems relevant to industries of national significance, such as in the mineral resources area. We seek to develop faster and more efficient separation technologies, and technologies for the manufacture, storage, and transport of particles.
Centre for Environmental Remediation
The Global Centre for Environmental Remediation (GCER) aims to safeguard people's social, economic and physical health and wellbeing by developing innovative, cost-effective and sustainable technologies and solutions that reduce the impact of pollutants on the environment.
Centre for Geotechnical Science and Engineering
The Centre for Geotechnical Science and Engineering develops new models and innovative computational methods for predicting the behaviour of geomaterials, metals and composites. Advanced computational methods, coupled with laboratory and field testing are key tools in this pursuit.
Centre for Innovative Energy Technologies
The Centre for Innovative Energy Technologies conducts cutting edge research on emerging energy technologies, with particular focus on the abatement of greenhouse gases, and clean and sustainable energy production.
Centre for Organic Electronics
The Centre for Organic Electronics is focused on the scientific challenges in the development of organic photovoltaics for the next generation of environmentally friendly energy sources, photonics and biosensors.
Centre for Secure and Reliable Communications
The Centre for Secure and Reliable Communications contributes to developing telecommunication networks that are robust to eavesdropping attacks, interference, and noise.
Centre for Space Science and Technology for Australian Resilience
Our Centre is transforming our understanding of the conditions in space. We use satellite technologies to observe our Sun and changing planet and support better ways to manage it. We also help shape policies to ensure space is used safely, responsibly, and for the benefit of all.
Innovative Centre for Advanced Nanomaterials
Innovation in materials science and research to develop advanced technologies and solutions for the global energy, environment and health sectors.
College of Health, Medicine and Wellbeing
Central Coast Research Institute
The Central Coast Research Institute (CCRI) is a joint venture of the University of Newcastle and Central Coast Local Health District. The CCRI aims to deliver pioneering research relevant to improving the health and wellbeing of the Central Coast community and beyond, focusing on the design, implementation and evaluation of new models of person-centred integrated care.
Centre for Drug Repurposing and Medicines Research
The Centre for Drug Repurposing & Medicines Research is committed to improve quality, safety, efficacy and timelines for bringing the most effective drug therapies to patients.
Centre for Gynaecological Diseases
The Centre for Gynaecological Diseases is a world-leading research centre focused on improving the gynaecological health of women and aims to pioneer a personalised approach to the prevention, detection, and treatment of gynaecological diseases.
Centre for Prevention, Implementation and Population Health
Centre for Prevention, Implementation and Population Health uses intervention and service delivery to promote healthy behaviours and good quality healthcare across communities.
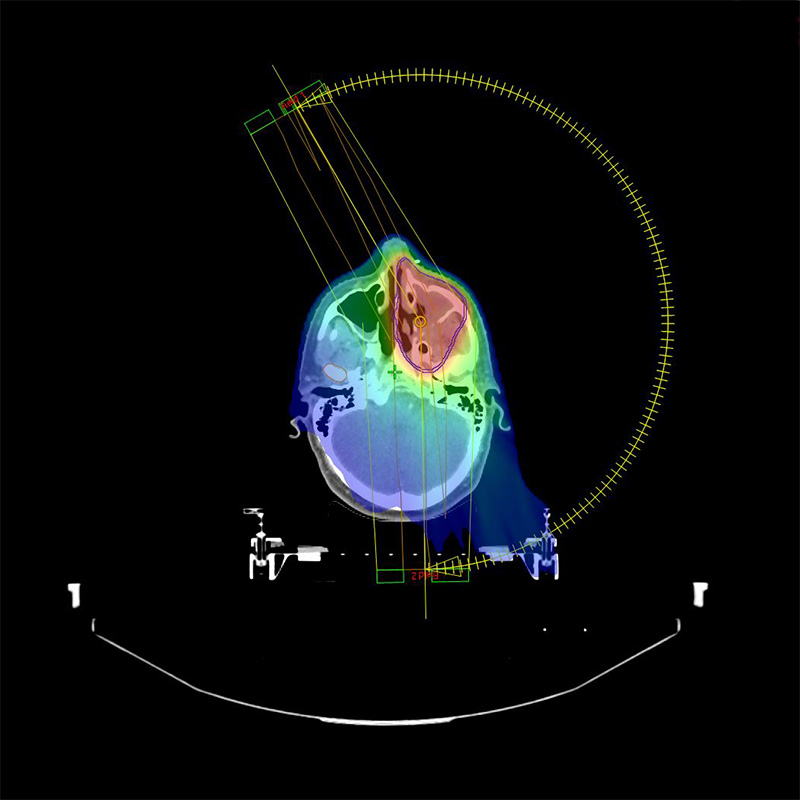
Centre for Research and Training in Radiation Oncology
The Centre is the first of its kind. With the support of industry partners and collaborators, the centre delivers education, research and training to domestic and international clinicians and students.
Centre for Women's Health Research
The Centre for Women's Health Research brings together academics and clinicians working in women’s health research and consumers with a passion for women’s health to research the factors that affect the health and wellbeing of women across the life course.
Global Sport and Movement Collaborative
The Global Sport and Movement Collaborative brings together world-class researchers, educators, and industry leaders to transform the future of sport, movement, and wellbeing.
Mark Hughes Foundation Centre for Brain Cancer Research
The Mark Hughes Foundation Centre for Brain Cancer Research is committed to finding a cure and improving the lives of those affected by brain cancer. We aim to advance brain cancer research and achieve the greatest impact for brain cancer patients and their families.
Medical Technology Research Centre
We pursue translational innovation in medical technology. We bridge disciplines across engineering, science and health, together with industry and clinical partners. Our mission is to engineer innovative medical devices to advance future healthcare.
College of Human and Social Futures
PhD scholarships and opportunities
Find out more
The University of Newcastle acknowledges the traditional custodians of the lands within our footprint areas: Awabakal, Darkinjung, Biripai, Worimi, Wonnarua, and Eora Nations. We also pay respect to the wisdom of our Elders past and present.